The patchwork chromosome. The human genome is a puzzle of duplications, duplications-within-duplications and more complex rearrangements.. Some of these duplications can misalign at meiosis and generate microdeletions and microduplications. The duplication architecture of the human genome is more pronounced in some chromosomes than in others. Chromosomes 15 and 16 are particularly rich in duplications, which is the reason several syndrome-associated microdeletions and microduplications are found there. One of these microdeletions is the 16p13.11 microdeletion. As a recent paper has looked as histological findings in brain tissue of patients with these deletions, it is time to review the only established genetic risk factor that contributes to wide range of epilepsy syndromes.
The 16p13.11 story. Hannes and colleagues first identified microdeletions in the 16p13.11 in several patients with multiple congenital anomalies. Even though most patients with 16p13.11 microdeletions do not have congenital anomalies, 16p13.11 joins the ranks of genetic risk factors that were first identified in extreme phenotypes. After the initial discovery, 16p13.11 was described in 0.5% of patients with IGE/GGE and 0.5% of patients with a wide range of epilepsy phenotypes including Temporal Lobe Epilepsy (TLE). Frequencies in schizophrenia and autism are likely to be comparable. There is some discrepancy regarding the frequency in the general population, but a reasonable estimate is 0.1%. This suggests that the risk for epilepsy in deletion carriers is increased by the factor of five. While this might sound impressive at first glance, the penetrance of this variant is actually quite low. Vassos and colleagues calculated the penetrance of many microdeletions associated with neurodevelopmental disorders and the results were quite surprising. Most microdeletions have a penetrance of 0.1 or less, suggesting that 90% or more of deletions carriers are unaffected. This estimate in combination with the increase in risk is perplexing. There is currently no consensus on how to handle this phenomenon in clinical practice let alone guidelines for counselling patients.

The 16p13.11 microdeletion. An 800kB region is flanked by segmental duplications, which may give rise to non-allelic homologous recombination, resulting in the loss or duplication of the intervening sequence.
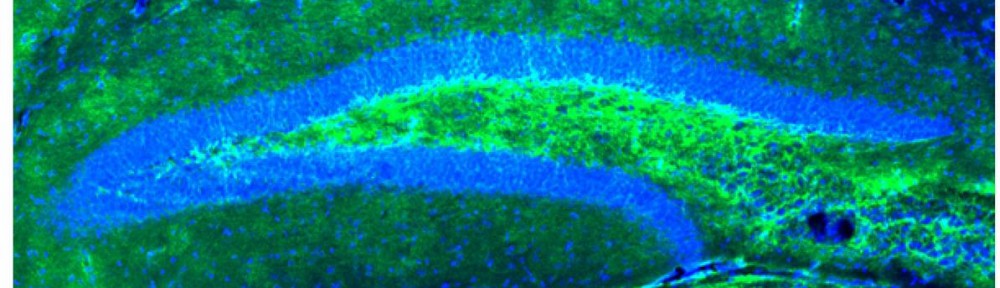
Neuropathology of 16p13.11. As some patients with 16p13.11 microdeletions have TLE, some patients underwent surgical removal of the temporal lobe. This gave Liu and colleagues the unique opportunity to examine the histology of the removed tissue from two patients. The results are quite simple: there is nothing extraordinary to see. One patient had a small hamartia (a small island of gray matter within the white matter), the other patient had histological findings of mesial temporal sclerosis (MTS), also referred to as hippocampal sclerosis. Both are common findings in patients with TLE. One might have expected subtle abnormal histological changes reflecting the underlying genetic defect, but the brain tissue was indistinguishable from the brain tissue resected from other patients. Liu and colleagues suggest that the epilepsy in 16p13.11 is not due to subtle histological defects.
The PLoS One effect. The Liu paper would not have been possible without PLoS One. PloS One is an open access journal dedicated to publishing all original research that is technically sound. No paper is rejected due to the lack of impact, but the impact of a paper will be judged by the responses (“post-publication impact”) to the published papers using citation metrics. It is difficult to publish negative findings and choosing PLOS One has provided the authors with an opportunity to simply present the results the way they are without trying to “sell” a larger story. The negative histological finding is one of the short stories that frequently remain unpublished. PLOS One now made small negative studies publishable and referenceable for the scientific community.
There is little, little to see in the brains of patients with 16p13. This little sentence should help you remember how to pronounce cytogenetic coordinates correctly, as this is a rhyme (“one-three” instead of “thirteen”). In summary, the 16p13.11 microdeletion is a rare genetic risk factor for a broad range of epilepsies. Brain tissue samples from patients with this microdeletion do not show specific patterns.
Pingback: Standing on the shoulders of giants: the EPICURE GWAS on Idiopathic Generalized Epilepsy | EuroEPINOMICS
Pingback: Double Impact | Beyond the Ion Channel
Pingback: 15q11.2 – the microdeletion spectre | Beyond the Ion Channel
Pingback: Face to face – atypical face shape and CNVs in epilepsy | Beyond the Ion Channel
Pingback: Close encounters of the third kind – rare genetic variants in families | Beyond the Ion Channel
Pingback: The 16p11.2 microdeletion: assessing the phenotypic range | Beyond the Ion Channel
Pingback: Dysregulation of a microRNA in the 22q11.2 microdeletion | Beyond the Ion Channel
Pingback: The tale of two planets: the expanding spectrum of STXBP1 | Beyond the Ion Channel
Pingback: “Meta-channelopathies” – RBFOX1 deletions and human epilepsy | Beyond the Ion Channel
Pingback: 16p13.11 microdeletions and the male bias | Beyond the Ion Channel
Pingback: Guilt by association: SCN1A in Temporal Lobe Epilepsy | Beyond the Ion Channel
Pingback: Identifying core phenotypes – epilepsy, ID and recurrent microdeletions | Beyond the Ion Channel
Pingback: 2013 in review: top three lists and the gene finding of the year | Beyond the Ion Channel
Pingback: The Channelopathist has left the building – here are our top ten posts of the last two years | Beyond the Ion Channel